Fusion Sense was conceived in response to one of the major limitations of modern digital health: the absence of context. In today’s health-tech ecosystem, patients are often analyzed in isolation, with their health status reduced to disconnected physiological metrics, detached from the environment in which they live.
This project proposes a more ambitious and realistic vision: human health is a dynamic system that continuously interacts with its surroundings.
It is not enough to monitor heart rate without considering how it is influenced by poor ventilation. Measuring oxygen saturation alone is insufficient without understanding how it reacts to increased humidity or degraded air quality. Fusion Sense bridges the gap between clinical and environmental data to build a context-aware, continuous, and real-time health model.
From Isolated Data to Contextual IntelligenceThe system integrates critical biomedical signals — ECG, SpO₂, and PPG — with environmental variables such as temperature, humidity, and air quality.
Instead of presenting these as independent data streams, Fusion Sense merges them into a unified intelligence layer capable of identifying cause–effect relationships between environmental conditions and physiological responses.
The ultimate goal is to demonstrate that by combining advanced sensing technologies with Edge AI, homes, offices, and shared spaces can be transformed into proactive health-aware environments — operating invisibly, efficiently, and with full respect for privacy.
Hardware Architecture: A Distributed e-Health ClusterTo realize this vision, the system is built upon a distributed mini-cluster of Arduino UNO Q boards, specifically designed to emulate a real-world edge computing environment.
The choice of the Arduino UNO Q is deliberate. Its high-performance Qualcomm SoC, combined with optimized power consumption, enables on-device execution of AI models. Furthermore, its high-precision ADC ensures reliable digitization of sensitive physiological signals — a critical requirement for biomedical systems.
The cluster consists of three specialized and decoupled nodes:
🔹 Master Node – Q-1Acts as the system orchestrator.It manages control logic, inter-node communication, and hosts an interactive Web-UI interface.
Through this interface, users can visualize biomedical signals in real time using dynamic graphs, enabling immediate interpretation of physiological status.
🔹 Classification Node – Q-2Dedicated exclusively to ECG signal analysis.This node runs trained classification models designed to detect patterns associated with potential cardiac abnormalities.
By isolating this task, the system ensures fast and stable inference even under computational load.
🔹 Fusion Node – Q-3The intelligent core of the system.Here, physiological signals and environmental variables converge to generate a predictive estimation of overall well-being, identifying trends and deviations that would remain hidden if each signal were analyzed independently.
Why This Architecture MattersThis distributed design:
- Prevents computational bottlenecks
- Improves scalability
- Enhances system robustness
- Reflects how real-world intelligent health systems could be deployed in homes or workplaces
Fusion Sense demonstrates that contextual health intelligence at the edge is not a future concept — it is technically achievable today.
The analytical core of Fusion Sense lies in the implementation of TinyML using Edge Impulse. The workflow begins with the acquisition and labeling of multimodal data (ECG, SpO₂, and PPG), which then passes through Digital Signal Processing (DSP) blocks to remove noise and extract physiologically relevant features.
Using the Impulse Designer, lightweight neural networks are trained and optimized specifically for resource-constrained hardware. These models are capable of identifying subtle correlations between environmental conditions and physiological responses — relationships that would be nearly impossible to detect through manual analysis.
Once optimized, the models are deployed directly onto the Arduino UNO Q boards, enabling fully local inference without reliance on cloud infrastructure. This results in:
- Millisecond-level latency
- Fully autonomous operation without internet connectivity
- Increased robustness and reliability in real-world scenarios
As previously described, the system is structured into three functional nodes, with Node 2 specifically responsible for cardiac pathology classification based on electrocardiographic (ECG) signals.
Dataset for Training and ValidationFor training and validating the machine learning models, we used the ECG Heartbeat Categorization Dataset, a dataset widely recognized in scientific literature.
This dataset consists of two collections of heartbeat signals derived from two benchmark ECG classification databases:
- MIT-BIH Arrhythmia Dataset
- PTB Diagnostic ECG Database
Both are publicly available through PhysioNet. The number of samples in both collections is sufficiently large to enable effective training of deep neural networks.
The dataset has been extensively used in studies involving deep learning architectures for heartbeat classification, as well as for evaluating transfer learning approaches in biomedical signal processing. The signals represent ECG beat morphologies under normal conditions, various types of arrhythmias, and myocardial infarction cases. All signals have been preprocessed and segmented such that each sample corresponds to a single heartbeat.
MIT-BIH Arrhythmia Dataset- Number of samples: 109, 446
- Number of categories: 5
- Sampling frequency: 125 Hz
- Data source: MIT-BIH Arrhythmia Dataset (PhysioNet)
- N (Normal): 0
- S (Supraventricular): 1
- V (Ventricular): 2
- F (Fusion): 3
- Q (Unknown): 4
- Number of samples: 14, 552
- Number of categories: 2
- Sampling frequency: 125 Hz
- Data source: PTB Diagnostic ECG Database (PhysioNet)
As an important methodological step, all signals were cropped, resampled, and zero-padded when necessary so that each sample has a fixed dimension of 188 data points. This uniform dimensionality facilitates direct integration into machine learning pipelines.
Data FormatThe dataset is distributed as a set of CSV files, where:
- Each row represents a single example (one heartbeat).
- The final value in each row corresponds to the class label.
- The remaining values represent the preprocessed ECG signal samples.
Mohammad Kachuee, Shayan Fazeli, and Majid Sarrafzadeh.“ECG Heartbeat Classification: A Deep Transferable Representation.”arXiv preprint arXiv:1805.00794, 2018.
One of the most significant challenges in this dataset is the severe class imbalance, which represents a major obstacle when training cardiac pathology classification models. This imbalance can introduce bias during training, leading the model to predominantly predict the majority class while significantly reducing its ability to correctly identify minority classes—often those corresponding to clinically important pathologies.
To address this issue, a Generative Adversarial Network (GAN) was implemented to generate realistic synthetic ECG signals. These artificial signals are used to increase the number of samples in underrepresented classes, thereby balancing the dataset distribution. As a result, the classification model can learn more robust and discriminative representations, improving its overall performance and enhancing its ability to detect less frequent cardiac conditions (Figure 3).
Using this generative network, synthetic ECG signals corresponding to minority classes are produced, allowing the missing samples to be completed and the dataset distribution to be balanced. This step is essential to prevent the model from overfitting to the dominant class and to improve its generalization capability.
Once the dataset was balanced, the training process for the cardiac pathology classification model was carried out using both real and synthetic signals. The classifier’s performance was evaluated using multiple metrics, with the confusion matrix—shown in Figure 4—serving as one of the primary tools to assess classification quality and to analyze the distribution of correct predictions and misclassifications across the different classes.
The obtained metrics demonstrate a strong balance between discriminative capability and model robustness (Figure 5), particularly considering that this is a multi-class classification problem based on a dataset that was originally highly imbalanced.
Area Under the ROC Curve (AUC = 0.94)An AUC value close to 1 indicates that the model has an excellent ability to distinguish between classes. This result suggests that the classifier can correctly separate different types of heartbeats in most cases, which is especially important in preventive healthcare applications where minimizing false negatives is critical.
In practical terms, the model reliably identifies pathological patterns, even in the presence of physiological noise.
Weighted Precision (0.78)Weighted precision indicates that when the model predicts a specific class, it is correct approximately 78% of the time, taking into account the class distribution within the dataset.
This value reflects a significant reduction in bias toward the majority class, confirming that the use of GAN-generated synthetic signals positively contributed to balancing the dataset.
Weighted Recall (0.77)Recall measures the model’s ability to correctly detect actual heartbeat instances for each class. A value of 77% indicates that the classifier successfully identifies most relevant events, including those belonging to minority classes associated with cardiac pathologies.
From a clinical perspective, this is particularly important, as it demonstrates that the system does not overlook critical abnormal events, thereby reducing the risk of dangerous omissions.
Weighted F1-Score (0.77)The F1-score combines precision and recall into a single metric. The obtained value confirms that the model achieves a solid balance between correctly detecting pathologies and avoiding false alarms.
This balance is especially important in continuous monitoring systems, where an excessive number of false positives could lead to alert fatigue for users or caregivers.
This first part of the system is directly integrated into one of the three Arduino UNO Q boards, where the cardiac classification model runs locally. In this configuration, the device is capable of processing and analyzing the ECG signal in real time, performing inference directly on the edge (on-device) without transmitting data to the cloud.
This integration significantly reduces latency, enhances the privacy of sensitive biomedical data, and ensures autonomous operation of the system—even in environments with limited or unreliable connectivity.
Data FusionThis section describes the process and underlying motivation for integrating multiple biosignals within the Fusion Sense system. It is important to highlight that, in the scientific literature, it is uncommon to find datasets that simultaneously and synchronously include multiple biosignals such as ECG, PCG/FCG, and PPG. Acquiring these physiological signals together represents a significant challenge from both an experimental and instrumentation perspective.
To properly evaluate the impact of signal fusion on health state estimation, it was necessary to construct a dedicated dataset. To achieve this, three independent datasets—each focused on a specific biosignal—were integrated. These datasets were subsequently harmonized and temporally aligned to enable the data fusion experiment and simulate a realistic multimodal monitoring scenario.
This approach allows the system to model a real-world context in which different biomedical sensors provide complementary physiological information. By combining these signals, the system can extract richer and more contextual patterns, overcoming the inherent limitations of analyzing each biosignal independently.
However, this approach introduces a key challenge: since the signals originate from different datasets, there is no direct sample-to-sample correspondence between them. As a result, it is not possible to directly correlate the raw signals without first applying several essential preprocessing steps:
- Signal normalization, to ensure comparable magnitudes and scales across different biosignals.
- Temporal alignment, to guarantee coherence between equivalent physiological windows.
- Extraction of common physiological features, enabling each signal to be represented in a compact and comparable form.
Through this preprocessing pipeline, correlation is performed in the feature space rather than on the raw signals. This step is critical unless the signals are perfectly synchronized at acquisition time.
Las siguietes imagenes, son el resultado de la adecuacion de las señales de los tres datasets.
During this training phase, the initial step consisted of constructing a curated dataset composed exclusively of biosignals acquired from patients without diagnosed pathologies. This baseline dataset included synchronized ECG, PPG, and FCG signals, ensuring signal quality, proper labeling, and consistent sampling frequency. The objective of using only non-pathological data at this stage was to establish a robust reference model of normal physiological behavior, particularly for anomaly detection tasks.
Once the dataset was assembled, it was automatically preprocessed and reformatted to comply with Edge Impulse’s ingestion requirements. This included signal segmentation into fixed-legth windows, normalization, metadata structuring, and export into a compatible format for feature extraction and training within the Edge Impulse environment. After uploading the dataset, the impulse design was configured, including signal processing blocks (feature extraction) and the corresponding learning blocks (classification and anomaly detection models).
Upon completion of the training process, the optimized models were exported as Docker containers. Two independent deployment pipelines were then implemented:
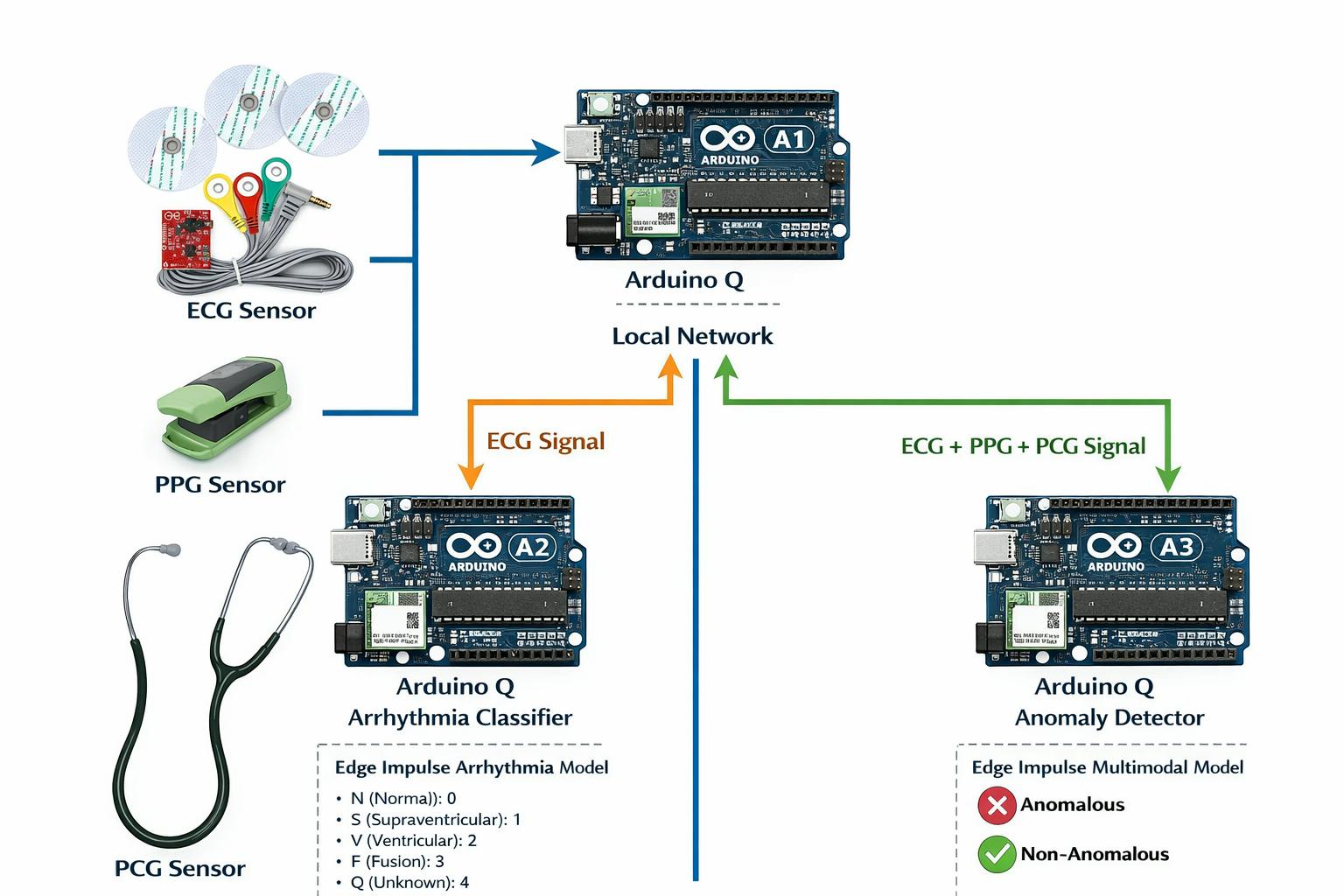
- The ECG pathology classifier was deployed on Arduino-Q2, where it performs real-time arrhythmia classification into the following categories:
- F (Fusion) = 0
- N (Normal) = 1
- Q (Unknown) = 2
- S (Supraventricular) = 3
- V (Ventricular) = 4
- The multimodal anomaly detection model was deployed on Arduino-Q3, where ECG, PPG, and FCG signals are fused to determine whether the combined physiological pattern corresponds to a normal or anomalous condition.
The three Arduino-Q nodes (Q1 for acquisition, Q2 for ECG classification, and Q3 for multimodal fusion analysis) were connected within the same local network to enable distributed edge inference. Each Docker container was executed independently on its respective device, allowing real-time processing, low-latency inference, and fully on-device decision-making without reliance on cloud-based computation.
The system was launched using the following command:
docker run --rm -it --network host public.ecr.aws/g7a8t7v6/inference-container-qc-xxxxxx-xxx:xx.xx.x --api-key <API-KEY> --run-http-server 1337Once the respective Docker containers are running on Arduino-Q2 and Arduino-Q3, Arduino-Q1 acts as the acquisition and orchestration node within the distributed edge architecture.
After capturing the ECG, PPG, and FCG signals, Arduino-Q1 formats the data into the predefined request structure and sends HTTP (or REST-based) inference requests over the local network to the deployed models:
- The ECG segment is transmitted to Arduino-Q2, where the arrhythmia classification model processes the signal and returns the predicted class (N, S, V, F, or Q) along with the corresponding confidence scores.
- The multimodal signal set (ECG + PPG + FCG) is sent to Arduino-Q3, where the anomaly detection model performs fusion-based analysis and determines whether the combined physiological pattern is classified as anomalous or non-anomalous.
Arduino-Q1 then receives the inference responses from both nodes, aggregates the results, and can optionally display them in the HTML interface, log them for further analysis, or trigger additional actions (e.g., alerts or status indicators).
This architecture enables fully distributed, real-time edge inference, where signal acquisition, classification, and anomaly detection are executed on separate devices within the same network, ensuring low latency, modularity, and independence from cloud-based processing.
Fusion Sense is more than a technological proof of concept—it is a scalable proposal for socially responsible healthcare innovation. The system has been designed with a clear objective: to democratize access to advanced physiological monitoring while preserving privacy and autonomy at the edge.
A central pillar of modern digital health is data sovereignty. Fusion Sense addresses this challenge by performing all signal processing, classification, and anomaly detection directly on-device. No biomedical data is transmitted to external cloud services, and no remote storage is required. This fully edge-based architecture eliminates the cybersecurity risks associated with centralized infrastructures, reduces exposure to data breaches, and ensures compliance with strict data protection standards. Patients retain control over their information, reinforcing trust in the technology.
Beyond privacy, the system is built around accessibility and prevention. Its modular design, low hardware cost, and autonomous operation make it suitable for deployment in resource-constrained environments. Because it does not depend on high-bandwidth connectivity or permanent cloud access, it can function reliably in remote or underserved regions.
Rural and Underserved CommunitiesFusion Sense enables preventive monitoring and early-stage risk detection in areas where access to specialized healthcare services is limited. By providing localized physiological analysis (ECG classification, multimodal anomaly detection), the system can support early triage decisions and reduce the need for long-distance medical travel. This is particularly relevant in geographically isolated communities, where delayed diagnosis often worsens health outcomes.
Elderly and Long-Term CareFor aging populations, continuous and non-invasive monitoring is essential. Fusion Sense supports long-term physiological supervision, allowing subtle deviations from baseline patterns to be detected before escalating into acute events. Early detection of arrhythmias or multimodal anomalies can contribute to proactive interventions, potentially reducing hospital admissions and improving quality of life.
Preventive and Occupational HealthThe architecture can also be adapted to workplace environments, supporting preventive health strategies in high-risk or physically demanding professions. By identifying abnormal physiological responses early, the system may help prevent incidents related to stress, fatigue, or cardiovascular instability.
Future WorkThe current version of the Multi-Sensor Health Monitor demonstrates real-time multimodal cardiac analysis (ECG, PPG and PCG) running entirely on-device using Arduino Q.
But this is just the beginning.
Robotic IntegrationA next step is connecting the system to a robotic platform capable of:
- Assisted sensor positioning
- Automated auscultation
- Remote interaction
- Adaptive signal acquisition
The robot would transform the project from a static monitor into an intelligent mobile health assistant.
Long-Range Communication with LoRaTo support rural and low-connectivity environments, the system will integrate:
- LoRa modules
- Low-power long-range transmission
- Distributed health alert broadcasting
This would allow multiple health nodes to communicate over kilometers without relying on traditional internet infrastructure.
Decentralized Networking with MeshtasticBy integrating Meshtastic:
- Devices could form autonomous mesh health networks
- Alerts could propagate peer-to-peer
- The system would become resilient in emergency or disaster scenarios
A fully decentralized, edge-based health monitoring ecosystem becomes possible.
Camera IntegrationFuture versions will include camera modules to enable:
- Visual confirmation of sensor placement
- Context-aware physiological analysis
- Multimodal AI fusion (vision + biosignals)
This opens the door to more intelligent and adaptive monitoring.
📶 4G Connectivity for Remote SupervisionFor critical events, the system will support optional 4G/LTE communication to:
- Send alerts to medical professionals
- Enable remote supervision
- Support hybrid Edge-Cloud architectures
Intelligence stays on-device, but critical situations can escalate safely.
Long-Term VisionThe goal is to evolve this project into:
- A distributed Edge-AI health ecosystem
- A robot-assisted biomedical platform
- A privacy-preserving telemedicine tool
- A resilient solution for underserved communities
https://www.hackster.io/jarain78/transmission-of-biosignals-via-lora-using-the-meshtastic-net-f1e4ba
Part II: Biosignal Transmission over LoRa Meshtastic Networkhttps://www.hackster.io/jarain78/part-ii-biosignal-transmission-over-lora-meshtastic-network-83f5eb
Intelligent Health Care Beaconhttps://www.hackster.io/jarain78/intelligent-health-care-beacon-3d154d
ECG Classification is the Edge Impulse project used to develop and deploy a lightweight cardiac anomaly detection model. Here, ECG data is collected, processed, and used to design optimized impulses for TinyML. The trained model runs directly on embedded hardware for real-time, on-device inference.
Fusion Data is the multimodal processing module running on the Arduino UNO Q cluster. It combines ECG, PPG, and FCG signals into a unified physiological representation using feature-level fusion. The system performs real-time, on-device analysis to extract contextual health insights without relying on the cloud.
Colab NotebooksVideoImage Galery



















_3u05Tpwasz.png?auto=compress%2Cformat&w=40&h=40&fit=fillmax&bg=fff&dpr=2)
Comments